META-Farm: Difference between revisions
mNo edit summary |
|||
| (87 intermediate revisions by 6 users not shown) | |||
| Line 1: | Line 1: | ||
<languages /> | |||
<translate> | |||
=Overview= <!--T:133--> | |||
<!--T:134--> | |||
META (short for META-Farm) is a suite of scripts designed in SHARCNET to automate high-throughput computing, that is, running a large number of related calculations. This practice is sometimes called <i>farming</i>, <i>serial farming</i>, or <i>task farming</i>. META works on all Alliance national systems, and could also be used on other clusters which use the same setup (most importantly, which use the [https://slurm.schedmd.com/documentation.html Slurm scheduler]). | |||
<!--T:135--> | |||
In this article, we use the term <b>case</b> for one independent computation, which may involve the execution of a serial program, a parallel program, or a GPU-using program.<br> | |||
The term <b>job</b> is an invocation of the Slurm job scheduler, which may handle several cases. | |||
<!--T:136--> | |||
META has the following features: | |||
<!--T:137--> | |||
* two modes of operation: | |||
** SIMPLE mode, which handles one case per job, | |||
** META mode, which handles many cases per job, | |||
* dynamic workload balancing in META mode, | |||
* captures the exit status of all individual cases, | |||
* automatically resubmits all the cases which failed or never ran, | |||
* submits and independently operates multiple <i>farms</i> (groups of cases) on the same cluster, | |||
* can automatically run a post-processing job once all the cases have been processed successfully. | |||
== | <!--T:138--> | ||
* | Some technical requirements: | ||
* Load the meta-farm module | * For each farm, each case to be computed must be described as a separate line in a table.dat file. | ||
* You can run multiple farms independently, but each farm must have its own directory. | |||
<!--T:139--> | |||
In META mode, the number of actual jobs (called <i>metajobs</i>) submitted by the package is usually much smaller than the number of cases to process. Each metajob can process multiple lines (multiple cases) from table.dat. A collection of metajobs will read lines from table.dat, starting from the first line, in a serialized manner using the [https://linux.die.net/man/1/lockfile lockfile] mechanism to prevent a race condition. This ensures a good dynamic workload balance between metajobs, as metajobs which handle shorter cases will process more of them. | |||
<!--T:140--> | |||
Not all metajobs need to run in META mode. The first metajob to run will start processing lines from table.dat; if and when the second job starts, it joins the first one, and so on. If the runtime of an individual metajob is long enough, all the cases might be processed with just a single metajob. | |||
== META vs. GLOST == <!--T:141--> | |||
<!--T:142--> | |||
The META package has important advantages over other approaches like [[GLOST]] where farm processing is done by bundling up all the jobs into a large parallel (MPI) job: | |||
* As the scheduler has full flexibility to start individual metajobs when it wants, the queue wait time can be dramatically shorter with the META package than with GLOST. Consider a large farm where 1000 CPU cores need to be used for 3 days; | |||
** with GLOST, with a 1000-way MPI job, queue wait time can be weeks, so it'll be weeks before you see your very first result; | |||
** with META, some metajobs start to run and produce the first results within minutes. | |||
* At the end of the farm computations; | |||
** with GLOST, some MPI ranks will finish earlier and will sit idle until the very last—the slowest—MPI rank ends; | |||
** with META, there is no such waste at the end of the farm: individual metajobs exit earlier if they have no more workload to process. | |||
* GLOST and other similar packages do not support automated resubmission of the cases which failed or never ran. META has this feature, and it is very easy to use. | |||
== The META webinar == <!--T:143--> | |||
<!--T:144--> | |||
A webinar was recorded on October 6th, 2021 describing the META package. You can view it [https://youtu.be/GcYbaPClwGE here]. | |||
=Quick start= <!--T:145--> | |||
<!--T:146--> | |||
If you are impatient to start using META, just follow the steps listed below. However, it is highly recommended to also read the rest of the page. | |||
<!--T:147--> | |||
* Log into a cluster. | |||
* Load the <code>meta-farm</code> module. | |||
$ module load meta-farm | $ module load meta-farm | ||
* | * Choose a name for a farm directory, e.g. <code>Farm_name</code>, and create it with the following command | ||
$ farm_init.run Farm_name | $ farm_init.run Farm_name | ||
* | * This will also create a few important files inside the farm directory, some of which you will need to customize. | ||
* Copy your | * Copy your executable and input files to the farm directory. (You may skip this step if you plan to use full paths everywhere.) | ||
* | * Edit the <code>table.dat</code> file inside the farm directory. This is a text file describing one case (one independent computation) per line. For examples, see one or more of | ||
* Modify | ** [[#single_case.sh|single_case.sh]] | ||
* Modify | ** [[META:_Advanced_features_and_troubleshooting#Example:_Numbered_input_files|Example: Numbered input files]] (advanced) | ||
** [[META:_Advanced_features_and_troubleshooting#Example:_Input_file_must_have_the_same_name|Example: Input file must have the same name]] (advanced) | |||
** [[META:_Advanced_features_and_troubleshooting#Using_all_the_columns_in_the_cases_table_explicitly|Using all the columns in the cases table explicitly]] (advanced) | |||
* Modify the <code>single_case.sh</code> script if needed. In many cases you don't have to make any changes. For more information see one or more of | |||
** [[#single_case.sh|single_case.sh]] | |||
** [[#STATUS_and_handling_errors|STATUS and handling errors]] | |||
** [[META:_Advanced_features_and_troubleshooting#Example:_Input_file_must_have_the_same_name|Example: Input file must have the same name]] (advanced) | |||
** [[META:_Advanced_features_and_troubleshooting#Using_all_the_columns_in_the_cases_table_explicitly|Using all the columns in the cases table explicitly]] (advanced) | |||
* Modify the <code>job_script.sh</code> file to suit your needs as described at [[#job_script.sh|job_script.sh]] below. In particular, use a correct account name, and set an appropriate job runtime. For more about runtimes, see [[#Estimating_the_runtime_and_number_of_metajobs|Estimating the runtime and number of metajobs]]. | |||
* Inside the farm directory, execute | * Inside the farm directory, execute | ||
$ submit.run -1 | $ submit.run -1 | ||
for the one case per job (SIMPLE) mode, or | for the one case per job (SIMPLE) mode, or | ||
$ submit.run N | $ submit.run N | ||
for the many cases per job (META) mode | for the many cases per job (META) mode, where <i>N</i> is the number of metajobs to use. <i>N</i> should be significantly smaller than the total number of cases. | ||
To run another farm concurrently with the first one, run | <!--T:148--> | ||
To run another farm concurrently with the first one, run <code>farm_init.run</code> again (providing a different farm name) and customize files <code>single_case.sh</code> and <code>job_script.sh</code> inside the new farm directory, then create a new table.dat file there. Also copy the executable and all the input files as needed. Now you can execute the <code>submit.run</code> command inside the second farm directory to submit the second farm. | |||
=List of commands= <!--T:149--> | |||
* <b>farm_init.run</b>: Initialize a farm. See [[#Quick start|Quick start]] above. | |||
* <b>submit.run</b>: Submit the farm to the scheduler. See [[#submit.run|submit.run]] below. | |||
* <b>resubmit.run</b>: Resubmit all computations which failed or never ran as a new farm. See [[#Resubmitting_failed_cases|Resubmitting failed cases]]. | |||
* <b>list.run</b>: List all the jobs with their current state for the farm. | |||
* <b>query.run</b>: Provide a short summary of the state of the farm, showing the number of queued, running, and completed jobs. More convenient than using <code>list.run</code> when the number of jobs is large. It will also print the progress—that is, the number of processed cases vs. the total number of cases—both for the current run, and globally. | |||
* <b>kill.run</b>: Kill all the running and queued jobs in the farm. | |||
* <b>prune.run</b>: Remove only queued jobs. | |||
* <b>Status.run</b>: (capital "S") List statuses of all processed cases. With the optional <code>-f</code>, the non-zero status lines (if any) will be listed at the end. | |||
* <b>clean.run</b>: Delete all the files in the farm directory (including subdirectories if any present), except for <code>job_script.sh, single_case.sh, final.sh, resubmit_script.sh, config.h,</code> and <code>table.dat</code>. It will also delete all files associated with this farm in the <code>/home/$USER/tmp</code> directory. Be very careful with this script! | |||
<!--T:150--> | |||
All of these commands (except for <code>farm_init.run</code> itself) have to be executed inside a farm directory, that is, a directory created by <code>farm_init.run</code>. | |||
=Small number of cases (SIMPLE mode)= <!--T:151--> | |||
<!--T:152--> | |||
Recall that a single execution of your code is a <b>case</b> and a <b>job</b> is an invocation of the Slurm scheduler. If: | |||
* the total number of cases is fairly small--- say, less than 500, and | |||
* each case runs for at least 20 minutes, | |||
then it is reasonable to dedicate a separate job to each case using the SIMPLE mode. | |||
Otherwise you should consider using the META mode to handle many cases per job, | |||
* | for which please see [[#Large number of cases (META mode)|Large number of cases (META mode)]] below. | ||
<!--T:153--> | |||
The three essential scripts are the command <code>submit.run</code>, and two user-customizable scripts <code>single_case.sh</code> and <code>job_script.sh</code>. | |||
= | ==submit.run== <!--T:154--> | ||
<i><b>Note:</b> This section applies to both SIMPLE and META modes.</i><br><br> | |||
The command <code>submit.run</code> has one obligatory argument, the number of jobs to submit, <i>N</i>: | |||
The | |||
<!--T:155--> | |||
<source lang="bash"> | <source lang="bash"> | ||
$ submit.run N [-auto] [optional_sbatch_arguments] | $ submit.run N [-auto] [optional_sbatch_arguments] | ||
</source> | </source> | ||
If N=-1, you are requesting the SIMPLE mode ( | <!--T:156--> | ||
If <i>N</i>=-1, you are requesting the SIMPLE mode (submit as many jobs as there are lines in table.dat). If <i>N</i> is a positive integer, you are requesting the META mode (multiple cases per job), with <i>N</i> being the number of metajobs requested. Any other value for <i>N</i> is not valid. | |||
<!--T:157--> | |||
If the optional switch <code>-auto</code> is present, the farm will resubmit itself automatically at the end, more than once if necessary, until all the cases from table.dat have been processed. This feature is described at [[META:_Advanced_features_and_troubleshooting#Resubmitting_failed_cases_automatically|Resubmitting failed cases automatically]]. | |||
If | <!--T:158--> | ||
If a file named <code>final.sh</code> is present in the farm directory, <code>submit.run</code> will treat it as a job script for a post-processing job and it will be launched automatically once all the cases from table.dat have been successfully processed. See [[META:_Advanced_features_and_troubleshooting#Running_a_post-processing_job_automatically|Running a post-processing job automatically]] for more details. | |||
If | <!--T:159--> | ||
If you supply any other arguments, they will be passed on to the Slurm command <code>sbatch</code> used to launch all metajobs for this farm. | |||
==single_case.sh== <!--T:160--> | |||
<i><b>Note:</b> This section applies to both SIMPLE and META modes.</i><br><br> | |||
<!--T:161--> | |||
The function of <code>single_case.sh</code> is to read one line from <code>table.dat</code>, parse it, and use the contents of that line to launch your code for one case. | |||
You may wish to customize <code>single_case.sh</code> for your purposes. | |||
<!--T:162--> | |||
The version of <code>single_case.sh</code> provided by <code>farm_init.run</code> treats each line in <code>table.dat</code> as a literal command and executes it in its own subdirectory <code>RUNyyy</code>, where <i>yyy</i> is the case number. Here is the relevant section of <code>single_case.sh</code>: | |||
<!--T:232--> | |||
<source lang="bash"> | <source lang="bash"> | ||
... | ... | ||
| Line 99: | Line 150: | ||
# $ID contains the case id from the original table (can be used to provide a unique seed to the code etc) | # $ID contains the case id from the original table (can be used to provide a unique seed to the code etc) | ||
# $COMM is the line corresponding to the case $ID in the original table, without the ID field | # $COMM is the line corresponding to the case $ID in the original table, without the ID field | ||
# $METAJOB_ID is the jobid for the current | # $METAJOB_ID is the jobid for the current metajob (convenient for creating per-job files) | ||
<!--T:233--> | |||
mkdir -p RUN$ID | mkdir -p RUN$ID | ||
cd RUN$ID | cd RUN$ID | ||
<!--T:234--> | |||
echo "Case $ID:" | echo "Case $ID:" | ||
<!--T:235--> | |||
# Executing the command (a line from table.dat) | # Executing the command (a line from table.dat) | ||
# It's allowed to use more than one shell command (separated by semicolons) on a single line | # It's allowed to use more than one shell command (separated by semicolons) on a single line | ||
eval "$COMM" | eval "$COMM" | ||
<!--T:236--> | |||
# Exit status of the code: | # Exit status of the code: | ||
STATUS=$? | STATUS=$? | ||
<!--T:237--> | |||
cd .. | cd .. | ||
# +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ | # +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ | ||
| Line 118: | Line 174: | ||
</source> | </source> | ||
<!--T:163--> | |||
Consequently, if you are using the unmodified <code>single_case.sh</code> then each line of <code>table.dat</code> should contain a complete command. | |||
This may be a compound command, that is, several commands separated by semicolons (;). | |||
<!--T:164--> | |||
Typically <code>table.dat</code> will contain a list of identical commands differentiated only by their arguments, but it need not be so. | |||
Any executable statement can go into <code>table.dat</code>. | |||
Your <code>table.dat</code> could look like this: | |||
</translate> | |||
/home/user/bin/code1 1.0 10 2.1 | /home/user/bin/code1 1.0 10 2.1 | ||
cp -f /input_dir/input1 .; /code_dir/code | cp -f ~/input_dir/input1 .; ~/code_dir/code | ||
./code2 < IC.2 | ./code2 < IC.2 | ||
<translate> | |||
<!--T:165--> | |||
If you intend to execute the same command for every case and don't wish to repeat it on every line of <code>table.dat</code>, | |||
then you can edit <code>single_case.sh</code> to include the common command. | |||
Then edit your <code>table.dat</code> to contain only the arguments and/or redirects for each case. | |||
<!--T:166--> | |||
For example, here is a modification of <code>single_case.sh</code> which includes the command | |||
(<code>/path/to/your/code</code>), takes the contents of <code>table.dat</code> as arguments to | |||
that command, and uses the case number <code>$ID</code> as an additional argument: | |||
* single_case.sh | <!--T:238--> | ||
* single_case.sh | |||
<source lang="bash"> | <source lang="bash"> | ||
... | |||
# ++++++++++++++++++++++ This part can be customized: ++++++++++++++++++++++++ | # ++++++++++++++++++++++ This part can be customized: ++++++++++++++++++++++++ | ||
# Here we use $ID (case number) as a unique seed for Monte-Carlo type serial farming: | # Here we use $ID (case number) as a unique seed for Monte-Carlo type serial farming: | ||
/path/to/your/code -par $COMM -seed $ID | /path/to/your/code -par $COMM -seed $ID | ||
STATUS=$? | |||
# +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ | |||
... | ... | ||
</source> | </source> | ||
* table.dat | * table.dat | ||
<source lang="bash"> | <source lang="bash"> | ||
12.56 | 12.56 | ||
| Line 148: | Line 218: | ||
</source> | </source> | ||
<!--T:167--> | |||
<b>Note 1:</b> If your code doesn't need to read any arguments from <code>table.dat</code>, you still have to generate <code>table.dat</code>, with the number of lines equal to the number of cases you want to compute. In this case, it doesn't matter what you put inside <code>table.dat</code>—all that matters is the total number of the lines. The key line in the above example might then look like | |||
<!--T:168--> | |||
/path/to/your/code -seed $ID | |||
<!--T:169--> | |||
<b>Note 2:</b> You do not need to insert line numbers at the beginning of each line of <code>table.dat</code>. The script <code>submit.run</code> will modify <code>table.dat</code> to add line numbers if it doesn't find them there. | |||
===STATUS and handling errors=== <!--T:170--> | |||
<!--T:171--> | |||
What is <code>STATUS</code> for in <code>single_case.sh</code>? It is a variable which should be set to “0” if your case was computed correctly, and some positive value (that is, greater than 0) otherwise. It is very important: It is used by <code>resubmit.run</code> to figure out which cases failed so they can be re-computed. In the provided version of <code>single_case.sh</code>, <code>STATUS</code> is set to the exit code of your program. This may not cover all potential problems, since some programs produce an exit code of zero even if something goes wrong. You can change how <code>STATUS</code> is set by editing <code>single_case.sh</code>. | |||
<!--T:172--> | |||
For example if your code is supposed to write a file (say, <code>out.dat</code>) at the end of each case, test whether the file exists and set <code>STATUS</code> appropriately. | |||
In the following code fragment, <code>$STATUS</code> will be positive if either the exit code from the program is positive, or if <code>out.dat</code> doesn't exist or is empty: | |||
</translate> | |||
<source lang="bash"> | <source lang="bash"> | ||
STATUS=$? | STATUS=$? | ||
| Line 166: | Line 244: | ||
fi | fi | ||
</source> | </source> | ||
<translate> | |||
==job_script.sh== <!--T:173--> | |||
<!--T:174--> | |||
The file job_script.sh is the | <i><b>Note:</b> This section applies to both SIMPLE and META modes.</i><br><br> | ||
The file <code>job_script.sh</code> is the job script which will be submitted to SLURM for all metajobs in your farm. | |||
Here is the default version created for you by <code>farm_init.run</code>: | |||
<!--T:230--> | |||
<source lang="bash"> | <source lang="bash"> | ||
#!/bin/bash | #!/bin/bash | ||
| Line 180: | Line 262: | ||
#SBATCH -A Your_account_name | #SBATCH -A Your_account_name | ||
<!--T:231--> | |||
# Don't change this line: | # Don't change this line: | ||
task.run | task.run | ||
</source> | </source> | ||
At the very least you | <!--T:175--> | ||
At the very least you should change the account name (the <code>-A</code> switch), and the metajob runtime (the <code>-t</code> switch). | |||
In SIMPLE mode, you should set the runtime to be somewhat longer than the longest expected individual case. | |||
'''Important:''' | <!--T:176--> | ||
'''Important:''' Your <code>job_script.sh</code> ''must'' include the runtime switch (either <code>-t</code> or <code>--time</code>). | |||
This cannot be passed to <code>sbatch</code> as an optional argument to <code>submit.run</code>. | |||
Sometimes the following problem happens: | <!--T:177--> | ||
Sometimes the following problem happens: A metajob may be allocated to a node which has a defect, thereby causing your program to fail instantly. For example, perhaps your program needs a GPU but the GPU you're assigned is malfunctioning, or perhaps the <code>/project</code> file system is not mounted. (Please report such a defective node to support@tech.alliancecan.ca if you detect one!) But when it happens, that single bad metajob can quickly churn through <code>table.dat</code>, so your whole farm fails. If you can anticipate such problems, you can add tests to <code>job_script.sh</code> before the <code>task.run</code> line. For example, the following modification will test for the presence of an NVidia GPU, and if none is found it will force the metajob to exit before it starts failing your cases: | |||
</translate> | |||
<source lang="bash"> | <source lang="bash"> | ||
nvidia-smi >/dev/null | |||
retVal=$? | retVal=$? | ||
if [ $retVal -ne 0 ]; then | if [ $retVal -ne 0 ]; then | ||
exit 1 | exit 1 | ||
fi | fi | ||
task.run | task.run | ||
</source> | </source> | ||
<translate> | |||
<!--T:178--> | |||
There is a utility <code>gpu_test</code> which does a similar job to <code>nvidia_smi</code> in the above example. | |||
On Graham, Cedar, or Beluga you can copy it to your <code>~/bin</code> directory: | |||
<!--T:179--> | |||
cp ~syam/bin/gpu_test ~/bin | |||
<!--T:180--> | |||
The META package has a built-in mechanism which tries to detect problems of this kind and kill a metajob which churns through the cases too quickly. The two relevant parameters, <code>N_failed_max</code> and <code>dt_failed</code> are set in the file <code>config.h</code>. The protection mechanism is triggered when the first <code>$N_failed_max</code> cases are very short - less than <code>$dt_failed</code> seconds in duration. The default values are 5 and 5, so by default a metajob will stop if the first 5 cases all finish in less than 5 seconds. If you get false triggering of this protective mechanism because some of your normal cases have runtime shorter than <code>$dt_failed</code>, reduce the value of <code>dt_failed</code> in <code>config.h</code>. | |||
==Output files== <!--T:181--> | |||
<i><b>Note:</b> This section applies to both SIMPLE and META modes.</i><br><br> | |||
<!--T:182--> | |||
Once one or more | Once one or more metajobs in your farm are running, the following files will be created in the farm directory: | ||
* OUTPUT/slurm- | * <code>OUTPUT/slurm-$JOBID.out</code>, one file per metajob containing its standard output, | ||
* STATUSES/status. | * <code>STATUSES/status.$JOBID</code>, one file per metajob containing the status of each case that was processed. | ||
In both cases, | <!--T:183--> | ||
In both cases, <code>$JOBID</code> stands for the jobid of the corresponding metajob. | |||
One more directory | <!--T:184--> | ||
One more directory, <code>MISC</code>, will also be created inside the root farm directory. It contains some auxiliary data. | |||
Also, every | <!--T:185--> | ||
Also, every time <code>submit.run</code> is run, it will create a unique subdirectory inside <code>/home/$USER/tmp</code>. | |||
Inside that subdirectory, some small scratch files will be created, such as files used by <code>lockfile</code> to serialize certain operations inside the jobs. | |||
These subdirectories have names <code>$NODE.$PID</code>, where <code>$NODE</code> is the name of the current node (typically a login node), and <code>$PID</code> is the unique process ID for the script. | |||
Once the farm execution is done, you can safely erase this subdirectory. | |||
This will happen automatically if you run <code>clean.run</code>, but be careful! <code>clean.run</code> also <b>deletes all the results</b> produced by your farm! | |||
==Resubmitting failed cases== <!--T:186--> | |||
<!--T:187--> | |||
The command | <i><b>Note:</b> This section applies to both SIMPLE and META modes.</i><br><br> | ||
The <code>resubmit.run</code> command takes the same arguments as <code>submit.run</code>: | |||
</translate> | |||
<source lang="bash"> | <source lang="bash"> | ||
$ resubmit.run N [-auto] [optional_sbatch_arguments] | $ resubmit.run N [-auto] [optional_sbatch_arguments] | ||
</source> | </source> | ||
<translate> | |||
<!--T:188--> | |||
<code>resubmit.run</code>: | |||
* analyzes all those <code>status.*</code> files (see [[#Output files|Output files]] above); | |||
* figures out which cases failed and which never ran for whatever reason (e.g. because of the metajobs' runtime limit); | |||
* creates or overwrites a secondary <code>table.dat_</code> file which lists only the cases that still need to be run; | |||
* launches a new farm for those cases. | |||
<!--T:189--> | |||
You cannot run <code>resubmit.run</code> until all the jobs from the original run are done or killed. | |||
<!--T:190--> | |||
If some cases still fail or do not run, you can resubmit the farm as many times as needed. Of course, if certain cases fail repeatedly, there must a be a problem with either the program you are running or its input. In this case you may wish to use the command <code>Status.run</code> (capital S!) which displays the statuses for all computed cases. With the optional argument <code>-f</code>, <code>Status.run</code> will sort the output according to the exit status, showing cases with non-zero status at the bottom to make them easier to spot. | |||
<!--T:191--> | |||
Similarly to <code>submit.run</code>, if the optional switch <code>-auto</code> is present, the farm will resubmit itself automatically at the end, more than once if necessary. This advanced feature is described at [[META:_Advanced_features_and_troubleshooting#Resubmitting_failed_cases_automatically|Resubmitting failed cases automatically]]. | |||
=Large number of cases (META mode)= <!--T:192--> | |||
The SIMPLE (one case per job) mode works fine when the number of cases is fairly small (<500). | |||
When the number of cases is much greater than 500, the following problems may arise: | |||
<!--T:193--> | |||
* Each cluster has a limit on how many jobs a user can have at one time. (e.g. for Graham, it is 1000.) | |||
* With a very large number of cases, each case computation is typically short. If one case runs for <20 min, CPU cycles may be wasted due to scheduling overheads. | |||
<!--T:194--> | |||
META mode is the solution to these problems. | |||
Instead of submitting a separate job for each case, a smaller number of <i>metajobs</i> are submitted, each of which processes multiple cases. | |||
To enable META mode the first argument to <code>submit.run</code> should be the desired number of metajobs, | |||
which should be a fairly small number—much smaller than the number of cases to process. e.g.: | |||
</translate> | |||
<source lang="bash"> | |||
$ submit.run 32 | |||
</source> | |||
<translate> | |||
This is how | <!--T:195--> | ||
Since each case may take a different amount of time to process, META mode uses a dynamic workload-balancing scheme. | |||
This is how META mode is implemented: | |||
<!--T:196--> | |||
[[File:meta1.png|500px]] | [[File:meta1.png|500px]] | ||
As the above diagram shows, | <!--T:197--> | ||
As the above diagram shows, each job executes the same script, <code>task.run</code>. Inside that script, there is a <code>while</code> loop for the cases. Each iteration of the loop has to go through a serialized portion of the code (that is, only one <i>job</i> at a time can be in that section of code), where it gets the next case to process from <code>table.dat</code>. Then the script <code>single_case.sh</code> (see [[#single_case.sh|single_case.sh]]) is executed once for each case, which in turn calls the user code. | |||
This approach results in dynamic workload balancing achieved across all the running | <!--T:198--> | ||
This approach results in dynamic workload balancing achieved across all the running <i>metajobs</i> belonging to the same farm. The algorithm is illustrated by the diagram below: | |||
<!--T:199--> | |||
[[File:DWB_META.png|800px]] | [[File:DWB_META.png|800px]] | ||
This can be seen more clearly in [https://www.youtube.com/watch?v=GcYbaPClwGE&t=423s this animation] | <!--T:200--> | ||
This can be seen more clearly in [https://www.youtube.com/watch?v=GcYbaPClwGE&t=423s this animation] from the META webinar. | |||
<!--T:201--> | |||
The dynamic workload balancing results in all metajobs finishing around the same time, regardless of how different the runtimes are for individual cases, regardless of how fast CPUs are on different nodes, and regardless of when individual <i>metajobs</i> start. In addition, not all metajobs need to start running for all the cases to be processed, and if a metajob dies (e.g. due to a node crash), at most one case will be lost. The latter can be easily rectified with <code>resubmit.run</code>; see [[META-Farm#Resubmitting_failed_cases|Resubmitting failed cases]]. | |||
< | <!--T:202--> | ||
Not all of the requested metajobs will necessarily run, depending on how busy the cluster is. But as described above, in META mode you will eventually get all your results regardless of how many metajobs run, although you might need to use <code>resubmit.run</code> to complete a particularly large farm. | |||
</ | |||
==Estimating the runtime and number of metajobs== <!--T:203--> | |||
<!--T:204--> | |||
How | How should you figure out the optimum number of metajobs, and the runtime to be used in <code>job_script.sh</code>? | ||
First you need to figure out | <!--T:205--> | ||
First you need to figure out the average runtime for an individual case (a single line in table.dat). Supposing your application program is not parallel, allocate a single CPU core with [[Running_jobs#Interactive_jobs|<code>salloc</code>]], then execute <code>single_case.sh</code> there for a few different cases. Measure the total runtime and divide that by the number of cases you ran to get an estimate of the average case runtime. This can be done with a shell <code>for</code> loop: | |||
</translate> | |||
<source lang="bash"> | <source lang="bash"> | ||
$ N=10; time for ((i=1; i<=$N; i++)); do ./single_case.sh table.dat $i ; done | $ N=10; time for ((i=1; i<=$N; i++)); do ./single_case.sh table.dat $i ; done | ||
</source> | </source> | ||
<translate> | |||
<!--T:206--> | |||
Divide the "real" time output by the above command by <code>$N</code> to get the average case runtime estimate. Let's call it <i>dt_case</i>. | |||
<!--T:207--> | |||
Estimate the total CPU time needed to process the whole farm by multiplying <i>dt_case</i> by the number of cases, that is, the number of lines in <code>table.dat</code>. | |||
If this is in CPU-seconds, dividing that by 3600 gives you the total number of CPU-hours. | |||
Multiply that by something like 1.1 or 1.3 to have a bit of a safety margin. | |||
Now you can make a sensible choice for the runtime of | <!--T:208--> | ||
Now you can make a sensible choice for the runtime of metajobs, and that will also determine the number of metajobs needed to finish the whole farm. | |||
The runtime you choose should be significantly larger | <!--T:209--> | ||
The runtime you choose should be significantly larger than the average runtime of an individual case, ideally by a factor of 100 or more. | |||
It must definitely be larger than the longest runtime you expect for an individual case. | |||
On the other hand it should not be too large; say, no more than 3 days. | |||
The longer a job's runtime is, the longer it will usually wait to be scheduled. | |||
On Alliance general-purpose clusters, a good choice would be 12h or 24h due to [[Job_scheduling_policies#Time_limits|scheduling policies]]. | |||
Once you have settled on a runtime, divide the total number of CPU-hours by the runtime you have chosen (in hours) to get the required number of metajobs. | |||
Round up this number to the next integer. | |||
<!--T:210--> | |||
With the above choices, the queue wait time should be fairly small, and the throughput and efficiency of the farm should be fairly high. | With the above choices, the queue wait time should be fairly small, and the throughput and efficiency of the farm should be fairly high. | ||
Let's consider a specific example. | <!--T:211--> | ||
Let's consider a specific example. Suppose you ran the above <code>for</code> loop on a dedicated CPU obtained with <code>salloc</code>, and the output said the "real" time was 15m50s, which is 950 seconds. Divide that by the number of sample cases, 10, to find that the average time for an individual case is 95 seconds. Suppose also the total number of cases you have to process (the number of lines in <code>table.dat</code>) is 1000. The total CPU time required to compute all your cases is then<br> | |||
95 x 1000 = 95,000 CPU-seconds = 26.4 CPU-hours<br> | |||
Multiply that by a factor of 1.2 as a safety measure, to yield 31.7 CPU-hours. A runtime of 3 hours for your metajobs would work here, and should lead to good queue wait times. Edit the value of the <code>#SBATCH -t</code> in <code>job_script.sh</code> to be <code>3:00:00</code>. Now estimate how many metajobs you'll need to process all the cases.<br> | |||
N = 31.7 core-hours / 3 hours = 10.6<br> | |||
which rounded up to the next integer is 11. Then you can launch the farm by executing a single <code>submit.run 11</code>. | |||
<!--T:212--> | |||
If the number of jobs in the above analysis is larger than 1000, you have a particularly large farm. | |||
The maximum number of jobs which can be submitted on Graham and Beluga is 1000, so you won't be able to run the whole collection with a single command. | |||
The workaround would be to go through the following sequence of commands. | |||
Remember each command can only be executed after the previous farm has finished running: | |||
</translate> | |||
<source lang="bash"> | <source lang="bash"> | ||
$ submit.run 1000 | $ submit.run 1000 | ||
| Line 297: | Line 450: | ||
... | ... | ||
</source> | </source> | ||
<translate> | |||
<!--T:213--> | |||
If this seems rather tedious, consider using an advanced feature of the META package for such large farms: [[META:_Advanced_features_and_troubleshooting#Resubmitting_failed_cases_automatically|Resubmitting failed cases automatically]]. This will fully automate the farm resubmission steps. | |||
== | =Words of caution= <!--T:223--> | ||
< | <!--T:224--> | ||
Always start with a small test run to make sure everything works before submitting a large production run. You can test individual cases by reserving an interactive node with <code>salloc</code>, changing to the farm directory, and executing commands like <code>./single_case.sh table.dat 1</code>, <code>./single_case.sh table.dat 2</code>, etc. | |||
< | |||
</ | |||
</ | |||
< | |||
</ | |||
< | <!--T:225--> | ||
If your farm is particularly large (say >10,000 cases), you should spend extra effort to make sure it runs as efficiently as possible. In particular, minimize the number of files and/or directories created during execution. If possible, instruct your code to append to existing files (one per metajob; <b>do not mix results from different metajobs in a single output file!</b>) instead of creating a separate file for each case. Avoid creating a separate subdirectory for each case. (Yes, creating a separate subdirectory for each case is the default setup of this package, but that default was chosen for safety, not efficiency!) | |||
</ | |||
It | <!--T:226--> | ||
The following example is optimized for a very large number of cases. It assumes, for purposes of the example: | |||
* that your code accepts the output file name via a command line switch <code>-o</code>, | |||
* that the application opens the output file in <b>append</b> mode, that is, multiple runs will keep appending to the existing file, | |||
* that each line of <code>table.dat</code> provides the rest of the command line arguments for your code, | |||
* that multiple instances of your code can safely run concurrently inside the same directory, so there is no need to create a subdirectory for each case, | |||
* and that each run will not produce any files besides the output file. | |||
With this setup, even very large farms (hundreds of thousands or even millions of cases) should run efficiently, as there will be very few files created. | |||
< | <!--T:227--> | ||
<source lang="bash"> | <source lang="bash"> | ||
... | ... | ||
| Line 543: | Line 479: | ||
# $ID contains the case id from the original table (can be used to provide a unique seed to the code etc) | # $ID contains the case id from the original table (can be used to provide a unique seed to the code etc) | ||
# $COMM is the line corresponding to the case $ID in the original table, without the ID field | # $COMM is the line corresponding to the case $ID in the original table, without the ID field | ||
# $METAJOB_ID is the jobid for the current | # $METAJOB_ID is the jobid for the current metajob (convenient for creating per-job files) | ||
<!--T:228--> | |||
# Executing the command (a line from table.dat) | # Executing the command (a line from table.dat) | ||
/path/to/your/code $COMM -o output.$METAJOB_ID | /path/to/your/code $COMM -o output.$METAJOB_ID | ||
<!--T:229--> | |||
# Exit status of the code: | # Exit status of the code: | ||
STATUS=$? | STATUS=$? | ||
| Line 554: | Line 492: | ||
</source> | </source> | ||
= If more help is needed = | = If more help is needed = <!--T:219--> | ||
<!--T:220--> | |||
See [[META-Farm: Advanced features and troubleshooting]] for more detailed discussion of some features, and for troubleshooting suggestions. | |||
<!--T:221--> | |||
If you need more help, contact [[technical support]], mentioning the name of the package (META), and the name of the staff member who wrote the software (Sergey Mashchenko). | |||
== Glossary == <!--T:222--> | |||
* '''case''': One independent computation. The file <code>table.dat</code> should list one case per line. | |||
* '''farm / farming''' (verb): Running many jobs on a cluster which carry out independent (but related) computations, of the same kind. | |||
* '''farm''' (noun): The directory and files involved in running one instance of the package. | |||
* '''metajob''': A job which can process multiple cases (independent computations) from <code>table.dat</code>. | |||
* '''META mode''': The mode of operation of the package in which each job can process ''multiple'' cases from <code>table.dat</code>. | |||
* '''SIMPLE mode''': The mode of operation of the package in which each job will process only one case from <code>table.dat</code>. | |||
</translate> | |||
Latest revision as of 20:44, 21 March 2024
Overview[edit]
META (short for META-Farm) is a suite of scripts designed in SHARCNET to automate high-throughput computing, that is, running a large number of related calculations. This practice is sometimes called farming, serial farming, or task farming. META works on all Alliance national systems, and could also be used on other clusters which use the same setup (most importantly, which use the Slurm scheduler).
In this article, we use the term case for one independent computation, which may involve the execution of a serial program, a parallel program, or a GPU-using program.
The term job is an invocation of the Slurm job scheduler, which may handle several cases.
META has the following features:
- two modes of operation:
- SIMPLE mode, which handles one case per job,
- META mode, which handles many cases per job,
- dynamic workload balancing in META mode,
- captures the exit status of all individual cases,
- automatically resubmits all the cases which failed or never ran,
- submits and independently operates multiple farms (groups of cases) on the same cluster,
- can automatically run a post-processing job once all the cases have been processed successfully.
Some technical requirements:
- For each farm, each case to be computed must be described as a separate line in a table.dat file.
- You can run multiple farms independently, but each farm must have its own directory.
In META mode, the number of actual jobs (called metajobs) submitted by the package is usually much smaller than the number of cases to process. Each metajob can process multiple lines (multiple cases) from table.dat. A collection of metajobs will read lines from table.dat, starting from the first line, in a serialized manner using the lockfile mechanism to prevent a race condition. This ensures a good dynamic workload balance between metajobs, as metajobs which handle shorter cases will process more of them.
Not all metajobs need to run in META mode. The first metajob to run will start processing lines from table.dat; if and when the second job starts, it joins the first one, and so on. If the runtime of an individual metajob is long enough, all the cases might be processed with just a single metajob.
META vs. GLOST[edit]
The META package has important advantages over other approaches like GLOST where farm processing is done by bundling up all the jobs into a large parallel (MPI) job:
- As the scheduler has full flexibility to start individual metajobs when it wants, the queue wait time can be dramatically shorter with the META package than with GLOST. Consider a large farm where 1000 CPU cores need to be used for 3 days;
- with GLOST, with a 1000-way MPI job, queue wait time can be weeks, so it'll be weeks before you see your very first result;
- with META, some metajobs start to run and produce the first results within minutes.
- At the end of the farm computations;
- with GLOST, some MPI ranks will finish earlier and will sit idle until the very last—the slowest—MPI rank ends;
- with META, there is no such waste at the end of the farm: individual metajobs exit earlier if they have no more workload to process.
- GLOST and other similar packages do not support automated resubmission of the cases which failed or never ran. META has this feature, and it is very easy to use.
The META webinar[edit]
A webinar was recorded on October 6th, 2021 describing the META package. You can view it here.
Quick start[edit]
If you are impatient to start using META, just follow the steps listed below. However, it is highly recommended to also read the rest of the page.
- Log into a cluster.
- Load the
meta-farmmodule.
$ module load meta-farm
- Choose a name for a farm directory, e.g.
Farm_name, and create it with the following command
$ farm_init.run Farm_name
- This will also create a few important files inside the farm directory, some of which you will need to customize.
- Copy your executable and input files to the farm directory. (You may skip this step if you plan to use full paths everywhere.)
- Edit the
table.datfile inside the farm directory. This is a text file describing one case (one independent computation) per line. For examples, see one or more of- single_case.sh
- Example: Numbered input files (advanced)
- Example: Input file must have the same name (advanced)
- Using all the columns in the cases table explicitly (advanced)
- Modify the
single_case.shscript if needed. In many cases you don't have to make any changes. For more information see one or more of - Modify the
job_script.shfile to suit your needs as described at job_script.sh below. In particular, use a correct account name, and set an appropriate job runtime. For more about runtimes, see Estimating the runtime and number of metajobs. - Inside the farm directory, execute
$ submit.run -1
for the one case per job (SIMPLE) mode, or
$ submit.run N
for the many cases per job (META) mode, where N is the number of metajobs to use. N should be significantly smaller than the total number of cases.
To run another farm concurrently with the first one, run farm_init.run again (providing a different farm name) and customize files single_case.sh and job_script.sh inside the new farm directory, then create a new table.dat file there. Also copy the executable and all the input files as needed. Now you can execute the submit.run command inside the second farm directory to submit the second farm.
List of commands[edit]
- farm_init.run: Initialize a farm. See Quick start above.
- submit.run: Submit the farm to the scheduler. See submit.run below.
- resubmit.run: Resubmit all computations which failed or never ran as a new farm. See Resubmitting failed cases.
- list.run: List all the jobs with their current state for the farm.
- query.run: Provide a short summary of the state of the farm, showing the number of queued, running, and completed jobs. More convenient than using
list.runwhen the number of jobs is large. It will also print the progress—that is, the number of processed cases vs. the total number of cases—both for the current run, and globally. - kill.run: Kill all the running and queued jobs in the farm.
- prune.run: Remove only queued jobs.
- Status.run: (capital "S") List statuses of all processed cases. With the optional
-f, the non-zero status lines (if any) will be listed at the end. - clean.run: Delete all the files in the farm directory (including subdirectories if any present), except for
job_script.sh, single_case.sh, final.sh, resubmit_script.sh, config.h,andtable.dat. It will also delete all files associated with this farm in the/home/$USER/tmpdirectory. Be very careful with this script!
All of these commands (except for farm_init.run itself) have to be executed inside a farm directory, that is, a directory created by farm_init.run.
Small number of cases (SIMPLE mode)[edit]
Recall that a single execution of your code is a case and a job is an invocation of the Slurm scheduler. If:
- the total number of cases is fairly small--- say, less than 500, and
- each case runs for at least 20 minutes,
then it is reasonable to dedicate a separate job to each case using the SIMPLE mode. Otherwise you should consider using the META mode to handle many cases per job, for which please see Large number of cases (META mode) below.
The three essential scripts are the command submit.run, and two user-customizable scripts single_case.sh and job_script.sh.
submit.run[edit]
Note: This section applies to both SIMPLE and META modes.
The command submit.run has one obligatory argument, the number of jobs to submit, N:
$ submit.run N [-auto] [optional_sbatch_arguments]
If N=-1, you are requesting the SIMPLE mode (submit as many jobs as there are lines in table.dat). If N is a positive integer, you are requesting the META mode (multiple cases per job), with N being the number of metajobs requested. Any other value for N is not valid.
If the optional switch -auto is present, the farm will resubmit itself automatically at the end, more than once if necessary, until all the cases from table.dat have been processed. This feature is described at Resubmitting failed cases automatically.
If a file named final.sh is present in the farm directory, submit.run will treat it as a job script for a post-processing job and it will be launched automatically once all the cases from table.dat have been successfully processed. See Running a post-processing job automatically for more details.
If you supply any other arguments, they will be passed on to the Slurm command sbatch used to launch all metajobs for this farm.
single_case.sh[edit]
Note: This section applies to both SIMPLE and META modes.
The function of single_case.sh is to read one line from table.dat, parse it, and use the contents of that line to launch your code for one case.
You may wish to customize single_case.sh for your purposes.
The version of single_case.sh provided by farm_init.run treats each line in table.dat as a literal command and executes it in its own subdirectory RUNyyy, where yyy is the case number. Here is the relevant section of single_case.sh:
...
# ++++++++++++++++++++++ This part can be customized: ++++++++++++++++++++++++
# Here:
# $ID contains the case id from the original table (can be used to provide a unique seed to the code etc)
# $COMM is the line corresponding to the case $ID in the original table, without the ID field
# $METAJOB_ID is the jobid for the current metajob (convenient for creating per-job files)
mkdir -p RUN$ID
cd RUN$ID
echo "Case $ID:"
# Executing the command (a line from table.dat)
# It's allowed to use more than one shell command (separated by semicolons) on a single line
eval "$COMM"
# Exit status of the code:
STATUS=$?
cd ..
# +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
...
Consequently, if you are using the unmodified single_case.sh then each line of table.dat should contain a complete command.
This may be a compound command, that is, several commands separated by semicolons (;).
Typically table.dat will contain a list of identical commands differentiated only by their arguments, but it need not be so.
Any executable statement can go into table.dat.
Your table.dat could look like this:
/home/user/bin/code1 1.0 10 2.1 cp -f ~/input_dir/input1 .; ~/code_dir/code ./code2 < IC.2
If you intend to execute the same command for every case and don't wish to repeat it on every line of table.dat,
then you can edit single_case.sh to include the common command.
Then edit your table.dat to contain only the arguments and/or redirects for each case.
For example, here is a modification of single_case.sh which includes the command
(/path/to/your/code), takes the contents of table.dat as arguments to
that command, and uses the case number $ID as an additional argument:
- single_case.sh
...
# ++++++++++++++++++++++ This part can be customized: ++++++++++++++++++++++++
# Here we use $ID (case number) as a unique seed for Monte-Carlo type serial farming:
/path/to/your/code -par $COMM -seed $ID
STATUS=$?
# +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
...
- table.dat
12.56
21.35
...
Note 1: If your code doesn't need to read any arguments from table.dat, you still have to generate table.dat, with the number of lines equal to the number of cases you want to compute. In this case, it doesn't matter what you put inside table.dat—all that matters is the total number of the lines. The key line in the above example might then look like
/path/to/your/code -seed $ID
Note 2: You do not need to insert line numbers at the beginning of each line of table.dat. The script submit.run will modify table.dat to add line numbers if it doesn't find them there.
STATUS and handling errors[edit]
What is STATUS for in single_case.sh? It is a variable which should be set to “0” if your case was computed correctly, and some positive value (that is, greater than 0) otherwise. It is very important: It is used by resubmit.run to figure out which cases failed so they can be re-computed. In the provided version of single_case.sh, STATUS is set to the exit code of your program. This may not cover all potential problems, since some programs produce an exit code of zero even if something goes wrong. You can change how STATUS is set by editing single_case.sh.
For example if your code is supposed to write a file (say, out.dat) at the end of each case, test whether the file exists and set STATUS appropriately.
In the following code fragment, $STATUS will be positive if either the exit code from the program is positive, or if out.dat doesn't exist or is empty:
STATUS=$?
if test ! -s out.dat
then
STATUS=1
fi
job_script.sh[edit]
Note: This section applies to both SIMPLE and META modes.
The file job_script.sh is the job script which will be submitted to SLURM for all metajobs in your farm.
Here is the default version created for you by farm_init.run:
#!/bin/bash
# Here you should provide the sbatch arguments to be used in all jobs in this serial farm
# It has to contain the runtime switch (either -t or --time):
#SBATCH -t 0-00:10
#SBATCH --mem=4G
#SBATCH -A Your_account_name
# Don't change this line:
task.run
At the very least you should change the account name (the -A switch), and the metajob runtime (the -t switch).
In SIMPLE mode, you should set the runtime to be somewhat longer than the longest expected individual case.
Important: Your job_script.sh must include the runtime switch (either -t or --time).
This cannot be passed to sbatch as an optional argument to submit.run.
Sometimes the following problem happens: A metajob may be allocated to a node which has a defect, thereby causing your program to fail instantly. For example, perhaps your program needs a GPU but the GPU you're assigned is malfunctioning, or perhaps the /project file system is not mounted. (Please report such a defective node to support@tech.alliancecan.ca if you detect one!) But when it happens, that single bad metajob can quickly churn through table.dat, so your whole farm fails. If you can anticipate such problems, you can add tests to job_script.sh before the task.run line. For example, the following modification will test for the presence of an NVidia GPU, and if none is found it will force the metajob to exit before it starts failing your cases:
nvidia-smi >/dev/null
retVal=$?
if [ $retVal -ne 0 ]; then
exit 1
fi
task.run
There is a utility gpu_test which does a similar job to nvidia_smi in the above example.
On Graham, Cedar, or Beluga you can copy it to your ~/bin directory:
cp ~syam/bin/gpu_test ~/bin
The META package has a built-in mechanism which tries to detect problems of this kind and kill a metajob which churns through the cases too quickly. The two relevant parameters, N_failed_max and dt_failed are set in the file config.h. The protection mechanism is triggered when the first $N_failed_max cases are very short - less than $dt_failed seconds in duration. The default values are 5 and 5, so by default a metajob will stop if the first 5 cases all finish in less than 5 seconds. If you get false triggering of this protective mechanism because some of your normal cases have runtime shorter than $dt_failed, reduce the value of dt_failed in config.h.
Output files[edit]
Note: This section applies to both SIMPLE and META modes.
Once one or more metajobs in your farm are running, the following files will be created in the farm directory:
OUTPUT/slurm-$JOBID.out, one file per metajob containing its standard output,STATUSES/status.$JOBID, one file per metajob containing the status of each case that was processed.
In both cases, $JOBID stands for the jobid of the corresponding metajob.
One more directory, MISC, will also be created inside the root farm directory. It contains some auxiliary data.
Also, every time submit.run is run, it will create a unique subdirectory inside /home/$USER/tmp.
Inside that subdirectory, some small scratch files will be created, such as files used by lockfile to serialize certain operations inside the jobs.
These subdirectories have names $NODE.$PID, where $NODE is the name of the current node (typically a login node), and $PID is the unique process ID for the script.
Once the farm execution is done, you can safely erase this subdirectory.
This will happen automatically if you run clean.run, but be careful! clean.run also deletes all the results produced by your farm!
Resubmitting failed cases[edit]
Note: This section applies to both SIMPLE and META modes.
The resubmit.run command takes the same arguments as submit.run:
$ resubmit.run N [-auto] [optional_sbatch_arguments]
resubmit.run:
- analyzes all those
status.*files (see Output files above); - figures out which cases failed and which never ran for whatever reason (e.g. because of the metajobs' runtime limit);
- creates or overwrites a secondary
table.dat_file which lists only the cases that still need to be run; - launches a new farm for those cases.
You cannot run resubmit.run until all the jobs from the original run are done or killed.
If some cases still fail or do not run, you can resubmit the farm as many times as needed. Of course, if certain cases fail repeatedly, there must a be a problem with either the program you are running or its input. In this case you may wish to use the command Status.run (capital S!) which displays the statuses for all computed cases. With the optional argument -f, Status.run will sort the output according to the exit status, showing cases with non-zero status at the bottom to make them easier to spot.
Similarly to submit.run, if the optional switch -auto is present, the farm will resubmit itself automatically at the end, more than once if necessary. This advanced feature is described at Resubmitting failed cases automatically.
Large number of cases (META mode)[edit]
The SIMPLE (one case per job) mode works fine when the number of cases is fairly small (<500). When the number of cases is much greater than 500, the following problems may arise:
- Each cluster has a limit on how many jobs a user can have at one time. (e.g. for Graham, it is 1000.)
- With a very large number of cases, each case computation is typically short. If one case runs for <20 min, CPU cycles may be wasted due to scheduling overheads.
META mode is the solution to these problems.
Instead of submitting a separate job for each case, a smaller number of metajobs are submitted, each of which processes multiple cases.
To enable META mode the first argument to submit.run should be the desired number of metajobs,
which should be a fairly small number—much smaller than the number of cases to process. e.g.:
$ submit.run 32
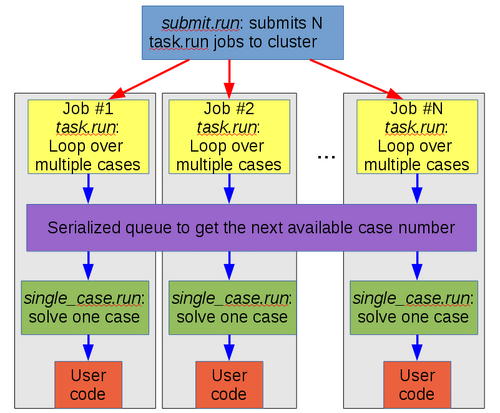
Since each case may take a different amount of time to process, META mode uses a dynamic workload-balancing scheme. This is how META mode is implemented:
As the above diagram shows, each job executes the same script, task.run. Inside that script, there is a while loop for the cases. Each iteration of the loop has to go through a serialized portion of the code (that is, only one job at a time can be in that section of code), where it gets the next case to process from table.dat. Then the script single_case.sh (see single_case.sh) is executed once for each case, which in turn calls the user code.
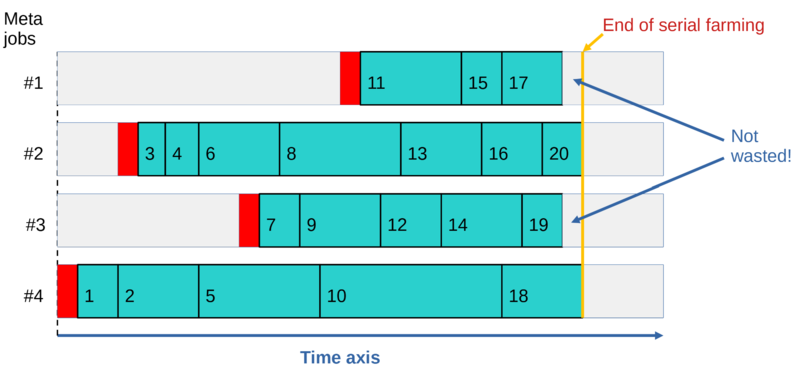
This approach results in dynamic workload balancing achieved across all the running metajobs belonging to the same farm. The algorithm is illustrated by the diagram below:
This can be seen more clearly in this animation from the META webinar.
The dynamic workload balancing results in all metajobs finishing around the same time, regardless of how different the runtimes are for individual cases, regardless of how fast CPUs are on different nodes, and regardless of when individual metajobs start. In addition, not all metajobs need to start running for all the cases to be processed, and if a metajob dies (e.g. due to a node crash), at most one case will be lost. The latter can be easily rectified with resubmit.run; see Resubmitting failed cases.
Not all of the requested metajobs will necessarily run, depending on how busy the cluster is. But as described above, in META mode you will eventually get all your results regardless of how many metajobs run, although you might need to use resubmit.run to complete a particularly large farm.
Estimating the runtime and number of metajobs[edit]
How should you figure out the optimum number of metajobs, and the runtime to be used in job_script.sh?
First you need to figure out the average runtime for an individual case (a single line in table.dat). Supposing your application program is not parallel, allocate a single CPU core with salloc, then execute single_case.sh there for a few different cases. Measure the total runtime and divide that by the number of cases you ran to get an estimate of the average case runtime. This can be done with a shell for loop:
$ N=10; time for ((i=1; i<=$N; i++)); do ./single_case.sh table.dat $i ; done
Divide the "real" time output by the above command by $N to get the average case runtime estimate. Let's call it dt_case.
Estimate the total CPU time needed to process the whole farm by multiplying dt_case by the number of cases, that is, the number of lines in table.dat.
If this is in CPU-seconds, dividing that by 3600 gives you the total number of CPU-hours.
Multiply that by something like 1.1 or 1.3 to have a bit of a safety margin.
Now you can make a sensible choice for the runtime of metajobs, and that will also determine the number of metajobs needed to finish the whole farm.
The runtime you choose should be significantly larger than the average runtime of an individual case, ideally by a factor of 100 or more. It must definitely be larger than the longest runtime you expect for an individual case. On the other hand it should not be too large; say, no more than 3 days. The longer a job's runtime is, the longer it will usually wait to be scheduled. On Alliance general-purpose clusters, a good choice would be 12h or 24h due to scheduling policies. Once you have settled on a runtime, divide the total number of CPU-hours by the runtime you have chosen (in hours) to get the required number of metajobs. Round up this number to the next integer.
With the above choices, the queue wait time should be fairly small, and the throughput and efficiency of the farm should be fairly high.
Let's consider a specific example. Suppose you ran the above for loop on a dedicated CPU obtained with salloc, and the output said the "real" time was 15m50s, which is 950 seconds. Divide that by the number of sample cases, 10, to find that the average time for an individual case is 95 seconds. Suppose also the total number of cases you have to process (the number of lines in table.dat) is 1000. The total CPU time required to compute all your cases is then
95 x 1000 = 95,000 CPU-seconds = 26.4 CPU-hours
Multiply that by a factor of 1.2 as a safety measure, to yield 31.7 CPU-hours. A runtime of 3 hours for your metajobs would work here, and should lead to good queue wait times. Edit the value of the #SBATCH -t in job_script.sh to be 3:00:00. Now estimate how many metajobs you'll need to process all the cases.
N = 31.7 core-hours / 3 hours = 10.6
which rounded up to the next integer is 11. Then you can launch the farm by executing a single submit.run 11.
If the number of jobs in the above analysis is larger than 1000, you have a particularly large farm. The maximum number of jobs which can be submitted on Graham and Beluga is 1000, so you won't be able to run the whole collection with a single command. The workaround would be to go through the following sequence of commands. Remember each command can only be executed after the previous farm has finished running:
$ submit.run 1000
$ resubmit.run 1000
$ resubmit.run 1000
...
If this seems rather tedious, consider using an advanced feature of the META package for such large farms: Resubmitting failed cases automatically. This will fully automate the farm resubmission steps.
Words of caution[edit]
Always start with a small test run to make sure everything works before submitting a large production run. You can test individual cases by reserving an interactive node with salloc, changing to the farm directory, and executing commands like ./single_case.sh table.dat 1, ./single_case.sh table.dat 2, etc.
If your farm is particularly large (say >10,000 cases), you should spend extra effort to make sure it runs as efficiently as possible. In particular, minimize the number of files and/or directories created during execution. If possible, instruct your code to append to existing files (one per metajob; do not mix results from different metajobs in a single output file!) instead of creating a separate file for each case. Avoid creating a separate subdirectory for each case. (Yes, creating a separate subdirectory for each case is the default setup of this package, but that default was chosen for safety, not efficiency!)
The following example is optimized for a very large number of cases. It assumes, for purposes of the example:
- that your code accepts the output file name via a command line switch
-o, - that the application opens the output file in append mode, that is, multiple runs will keep appending to the existing file,
- that each line of
table.datprovides the rest of the command line arguments for your code, - that multiple instances of your code can safely run concurrently inside the same directory, so there is no need to create a subdirectory for each case,
- and that each run will not produce any files besides the output file.
With this setup, even very large farms (hundreds of thousands or even millions of cases) should run efficiently, as there will be very few files created.
...
# ++++++++++++++++++++++ This part can be customized: ++++++++++++++++++++++++
# Here:
# $ID contains the case id from the original table (can be used to provide a unique seed to the code etc)
# $COMM is the line corresponding to the case $ID in the original table, without the ID field
# $METAJOB_ID is the jobid for the current metajob (convenient for creating per-job files)
# Executing the command (a line from table.dat)
/path/to/your/code $COMM -o output.$METAJOB_ID
# Exit status of the code:
STATUS=$?
# +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
...
If more help is needed[edit]
See META-Farm: Advanced features and troubleshooting for more detailed discussion of some features, and for troubleshooting suggestions.
If you need more help, contact technical support, mentioning the name of the package (META), and the name of the staff member who wrote the software (Sergey Mashchenko).
Glossary[edit]
- case: One independent computation. The file
table.datshould list one case per line. - farm / farming (verb): Running many jobs on a cluster which carry out independent (but related) computations, of the same kind.
- farm (noun): The directory and files involved in running one instance of the package.
- metajob: A job which can process multiple cases (independent computations) from
table.dat. - META mode: The mode of operation of the package in which each job can process multiple cases from
table.dat. - SIMPLE mode: The mode of operation of the package in which each job will process only one case from
table.dat.